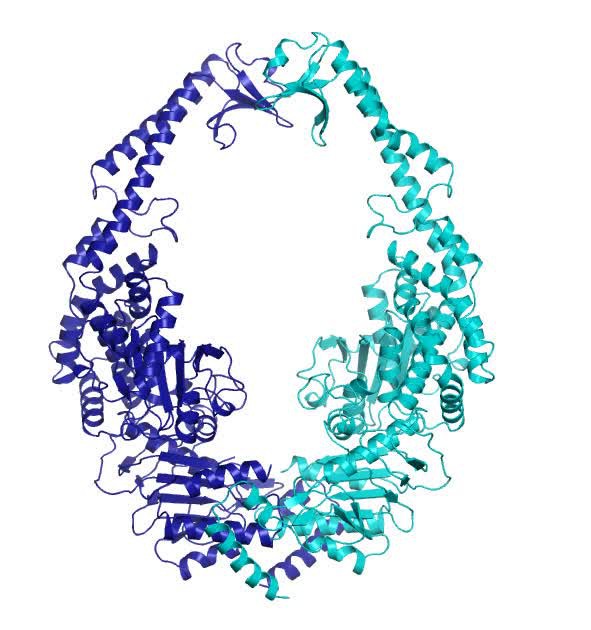
PDF) Using stable MutS dimers and tetramers to quantitatively analyze DNA mismatch recognition and sliding clamp formation

Pericentromeric chromatin reorganisation follows the initiation of recombination and coincides with early events of synapsis in cereals - Lenykó‐Thegze - 2021 - The Plant Journal - Wiley Online Library

Artificial surface labelling of Escherichia coli with StrepTagII antigen to study how monoclonal antibodies drive complement-mediated killing | Scientific Reports

DNA mismatch and damage detection using a FRET-based assay for monitoring the loading of multiple MutS sliding clamps | bioRxiv

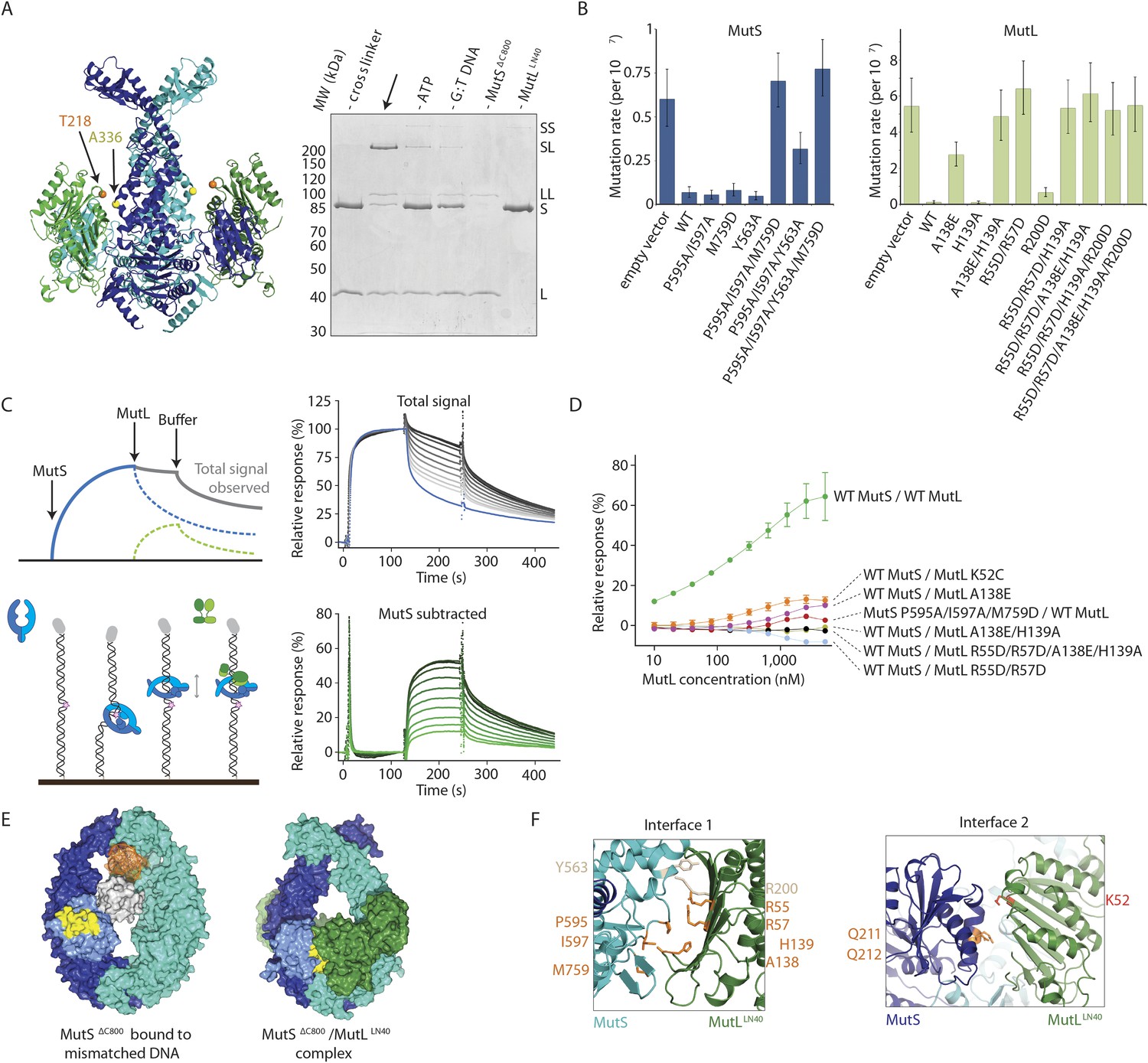
Cryo-EM structures reveal how ATP and DNA binding in MutS coordinate the sequential steps of DNA mismatch repair | bioRxiv
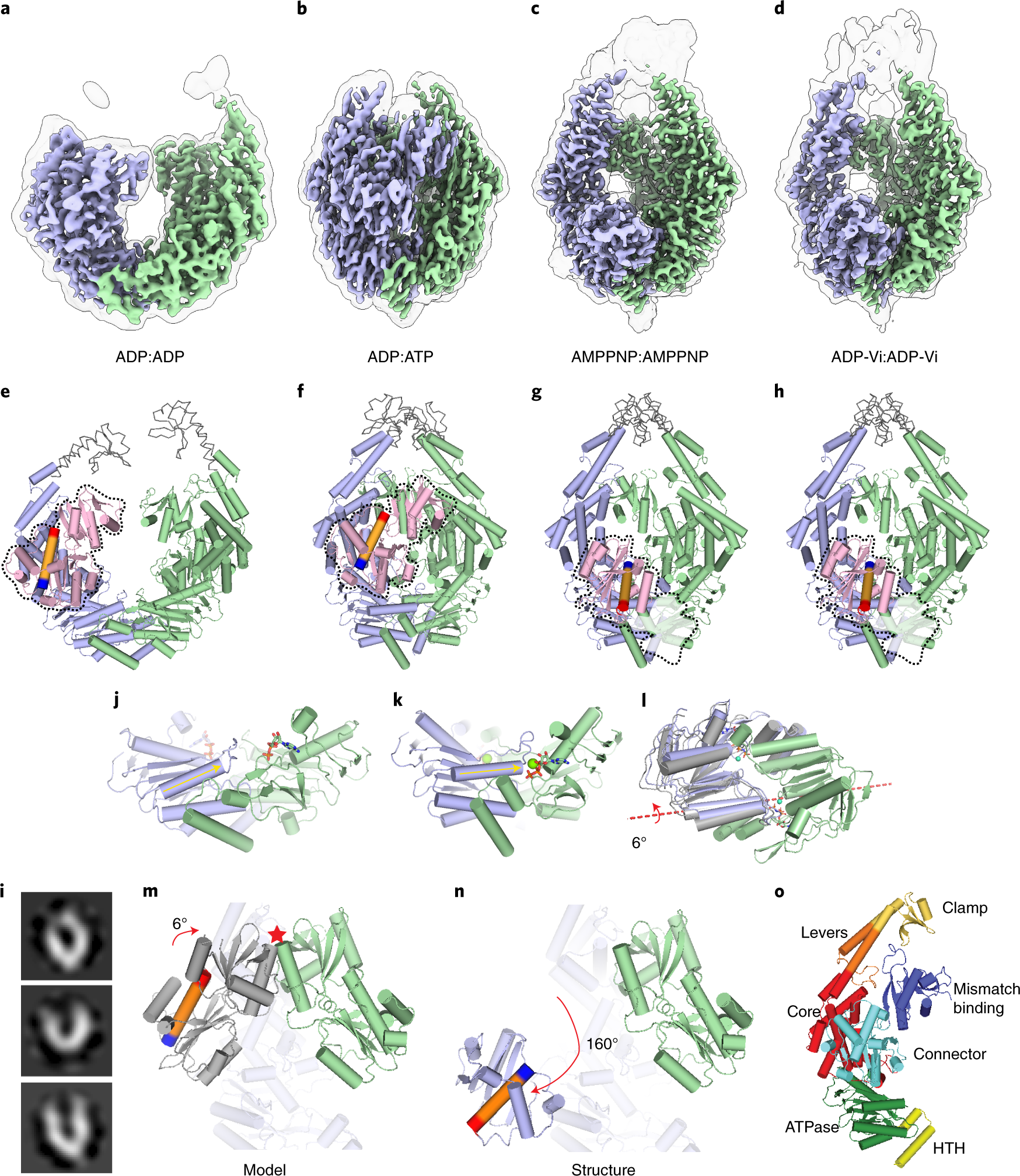
Cryogenic electron microscopy structures reveal how ATP and DNA binding in MutS coordinates sequential steps of DNA mismatch repair | Nature Structural & Molecular Biology

Mismatch Repair Inhibits Homeologous Recombination via Coordinated Directional Unwinding of Trapped DNA Structures - ScienceDirect

Unveiling the Inner Workings of Live Bacteria Using Super-Resolution Microscopy | Analytical Chemistry

PDF) Emergence of the Novel Aminoglycoside Acetyltransferase Variant aac(6′)-Ib-D179Y and Acquisition of Colistin Heteroresistance in Carbapenem-Resistant Klebsiella pneumoniae Due to a Disrupting Mutation in the DNA Repair Enzyme MutS

Cellular Recognition of Tri-/Di-palmitoylated Peptides Is Independent from a Domain Encompassing the N-terminal Seven Leucine-rich Repeat (LRR)/LRR-like Motifs of TLR2 - ScienceDirect

PDF) Using stable MutS dimers and tetramers to quantitatively analyze DNA mismatch recognition and sliding clamp formation

Mismatch Repair Inhibits Homeologous Recombination via Coordinated Directional Unwinding of Trapped DNA Structures - ScienceDirect
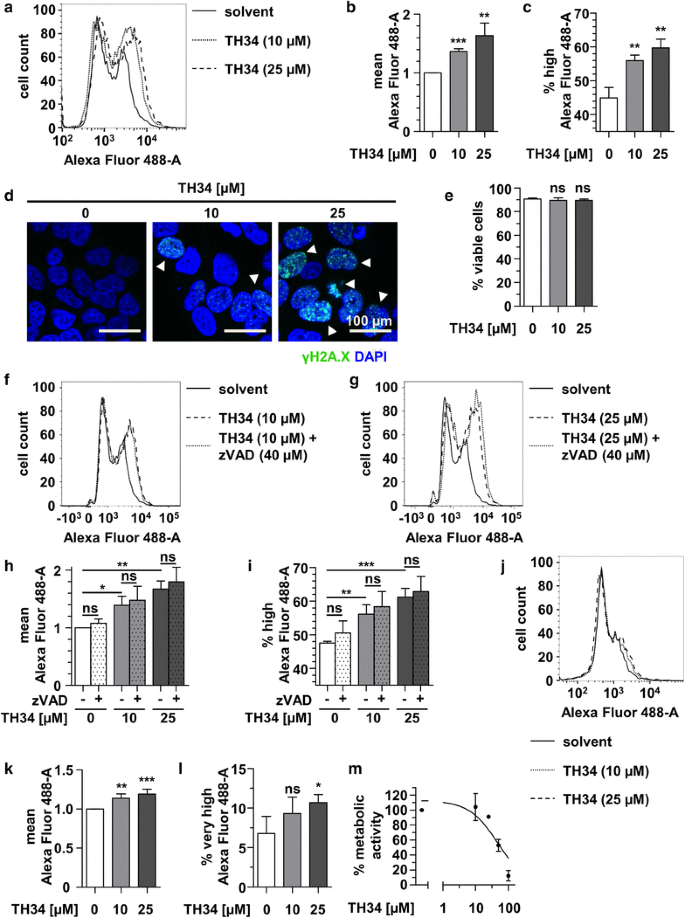
The HDAC6/8/10 inhibitor TH34 induces DNA damage-mediated cell death in human high-grade neuroblastoma cell lines | Archives of Toxicology

PDF) Using stable MutS dimers and tetramers to quantitatively analyze DNA mismatch recognition and sliding clamp formation

PDF) Outer membrane permeabilization by the membrane attack complex sensitizes Gram-negative bacteria to antimicrobial proteins in serum and phagocytes

Cryo-EM structures reveal how ATP and DNA binding in MutS coordinate the sequential steps of DNA mismatch repair | bioRxiv

Pericentromeric chromatin reorganisation follows the initiation of recombination and coincides with early events of synapsis in cereals - Lenykó‐Thegze - 2021 - The Plant Journal - Wiley Online Library

DNA mismatch and damage detection using a FRET-based assay for monitoring the loading of multiple MutS sliding clamps | bioRxiv



![Alexa Fluor® 488 Anti-MLH1 antibody [EPR3894] (ab199237) | Abcam Alexa Fluor® 488 Anti-MLH1 antibody [EPR3894] (ab199237) | Abcam](https://www.abcam.com/ps/products/199/ab199237/Images/ab199237-2-benefits-of-recombinant-antibodies.png)

![Recombinant Anti-MLH1 antibody [EPR3894] KO Tested (ab92312) | Abcam Recombinant Anti-MLH1 antibody [EPR3894] KO Tested (ab92312) | Abcam](https://www.abcam.com/ps/products/92/ab92312/Images/ab92312-16-anti-mlh1-antibody-epr3894-hap-1-flow-cytometry.jpg)

![Alexa Fluor® 488 Anti-MLH1 antibody [EPR3894] (ab199237) | Abcam Alexa Fluor® 488 Anti-MLH1 antibody [EPR3894] (ab199237) | Abcam](https://www.abcam.com/ps/products/199/ab199237/Images/ab199237-248777-anti-mlh1-antibody-epr3894-alexa-fluor-488-immunofluorescence.jpg)









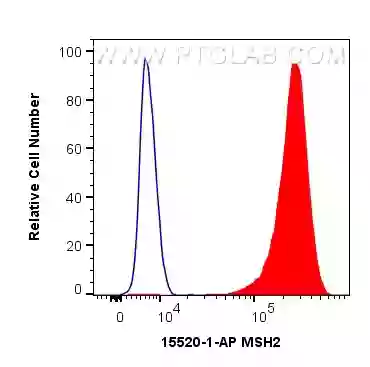
![Anti-MSH2 antibody [EPR21017-2] Recombinant (ab212188) | Abcam Anti-MSH2 antibody [EPR21017-2] Recombinant (ab212188) | Abcam](https://www.abcam.com/ps/products/212/ab212188/Images/ab212188-369004-anti-msh2-antibody-epr21017-2-msh2-ko-hap1-flow-cytometry-human.jpg)
![Recombinant Anti-MSH2 antibody [EPR21017-123] KO Tested (ab227941) | Abcam Recombinant Anti-MSH2 antibody [EPR21017-123] KO Tested (ab227941) | Abcam](https://www.abcam.com/ps/products/227/ab227941/Images/ab227941-291673-anti-msh2-antibody-epr21017-123-flow-cytometry.jpg)